HISTONES AND HMG-LIKE PROTEINS FROM PISUM SATIVUM CORRELATED WITH
EQUIVALENT FRACTIONS OF CALF THYMUS
Herlt, M. Institute of Genetics
University of Bonn, Federal Republic of Germany
Eucaryotic DNA is closely associated with a wide variety of DNA-
binding proteins. Traditionally, they are divided into two different
classes, the histones and the nonhistone chromosomal proteins. Due to
the basic nature of the histones, caused by the high proportion of posi-
tively charged amino acids such as lysine and arginine, these proteins
bind tightly to DNA nearly all the time and regardless of its nucleotide
sequence. Histones act as structural elements for folding the DNA into
nucleosomal strands. The fundamental packing unit, the nucleosome, is
formed by the core histones H2A, H2B, H3, and H4. H1 histones appear to
be responsible for organizing nucleosomes into regular higher-order
structures. The heterogeneity in histone H1 as well as the high rate of
post-translational modification events suggest a functional role by the
induction of conformational changes in the chromatin structure.
The abundant group of nonhistone proteins (NHP's) comprises the
remaining DNA-binding proteins. Due to their low concentration in the
nucleus, these putative gene regulatory proteins are very difficult to
isolate in sufficient amounts. A well defined class of NHP's is repre-
sented by the high mobility group or HMG proteins, so called because
they are relatively small and highly charged and, therefore, move
quickly during electrophoresis. HMG 14 and HMG 17 are characterized by
their specific interaction with nucleosomes associated with active
genes. At present participation of the HMG-proteins in gene regulation
is described only in animals. Investigations aimed at identifying HMG-
equivalents in plants are needed.
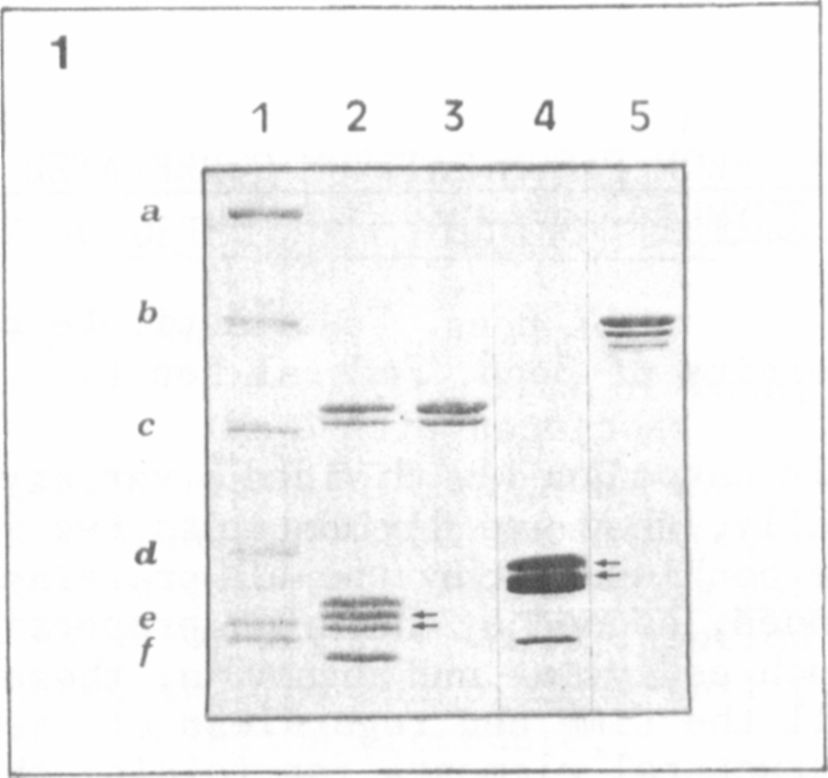
In this paper Fig. 1 shows an electropherogram of a 10-20% SDS-gel
(Laemmli-system) comparing purified histone fractions of calf thymus
with Pisum sativum. A difference can be seen in the electrophoretic
mobilities of the H2-group proteins (lane 2,4, arrows). This plant-
specific protein pattern of H2A/H2B (lane 4) is reconfirmed in compari-
sons of histone H1 complexes (purified by gel filtration). The two main
variants of calf thymus HI (lane 3) exhibit a relative molecular mass of
about 30 KD. In contrast, at least three pea HI bands are found at
about 40 KD.
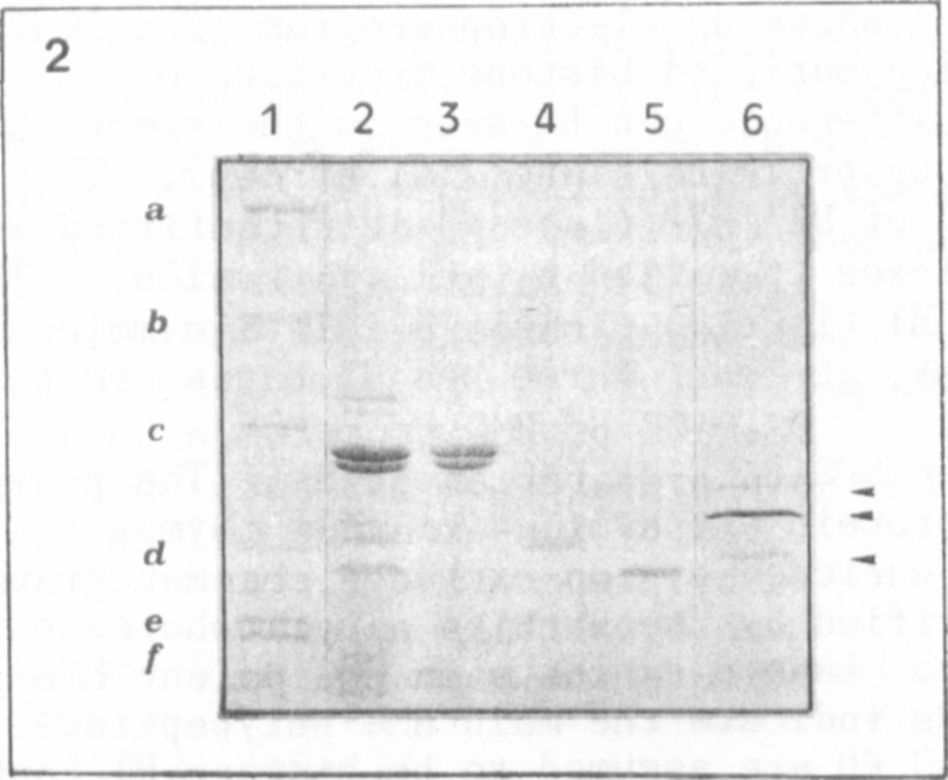
Fig. 2 represents an SDS-PAGE of HMG proteins. The polypeptide
pattern of a crude HMG protein preparation of calf thymus is shown in
lane 2, HMG 1 and HMG 2 enriched by ion-exchange chromatography in lane
3, HMG 14 and HMG 17 purified by preparative electrophoresis in lane 4
and lane 5, respectively. Lane 6 exhibits an equivalent HMG preparation
of Pisum. The arrowheads indicate the main HMG polypeptides. Faint
bands located at about 40 KD are assumed to be histone HI contamina-
tions.
The similarity in extractability and electrophoretic mobility of
mammalian HMG's and the corresponding pea protein fraction permits the
conclusion that HMG equivalents exist in plants. However, in the case
of HMG's as well as histone H2-group and H1 complex a plant specificity
can be supposed.